A new GBIF Plazi data issue feedback loop
Plazi’s goal is to discover known biodiversity, and make it widely available using the tools available in the digital age. This means liberating data hidden in libraries, and more recently, and increasingly hidden in the PDF-prison, paywalls and unstructured text.
The grand challenge is an estimated 500 Million published pages of scholarly biodiversity related publications, and 17,000 annually new discovered and described species with a multiple of annotations on already known species.
This clearly cannot be done by an individual institution, despite the rapidly developing technology. It needs collaboration and a strategy. With its rapidly growing corpus of liberated taxonomic treatments and technology, Plazi hopes to inspire others to join this endeavour. While the greatest interest and richness in the data lies in the details - who collected when where what species - this can only be achieved, if its context is liberated, that is accessible as FAIR taxonomic treatments.
Our current strategy is to make one million taxonomic treatments openly accessible, including taxonomic names, and by leveraging treatment citations, build the catalogue of life with each name linked to the scientific argument and data. We are well on our way.
To tackle this, tools and infrastructures are developed, if necessary. They allow converting PDF documents into a text and multimedia stream that can be further processed by adding tags enhancing single words to entire sections with a meaning (semantic enhancement) so humans and machines can understand its content. This is quite a technical challenge.
Materials citations attract great attention. They reference specimens in various formats used in the research process leading to the research results - in the biodiversity world - the published taxonomic treatments. However, they are a collateral in the current production. We try to discover them, make them accessible via the Global Biodiversity Information Facility (GBIF) as occurrences. We routinely quality control holotype citations whether they are properly extracted, and let the others pass, unless dedicated resources are available.
We believe that, despite this raw form, we provide a services by drawing attention to specimens in collections and species that are not recorded in GBIF, and that this will eventually lead to better data. It will raise the awareness of authors and publishers of the value to publish specimen data in a standardized way, as proposed by EJT and Pensoft.
We also believe that together with your input, we can make the material citations fitter for more uses.
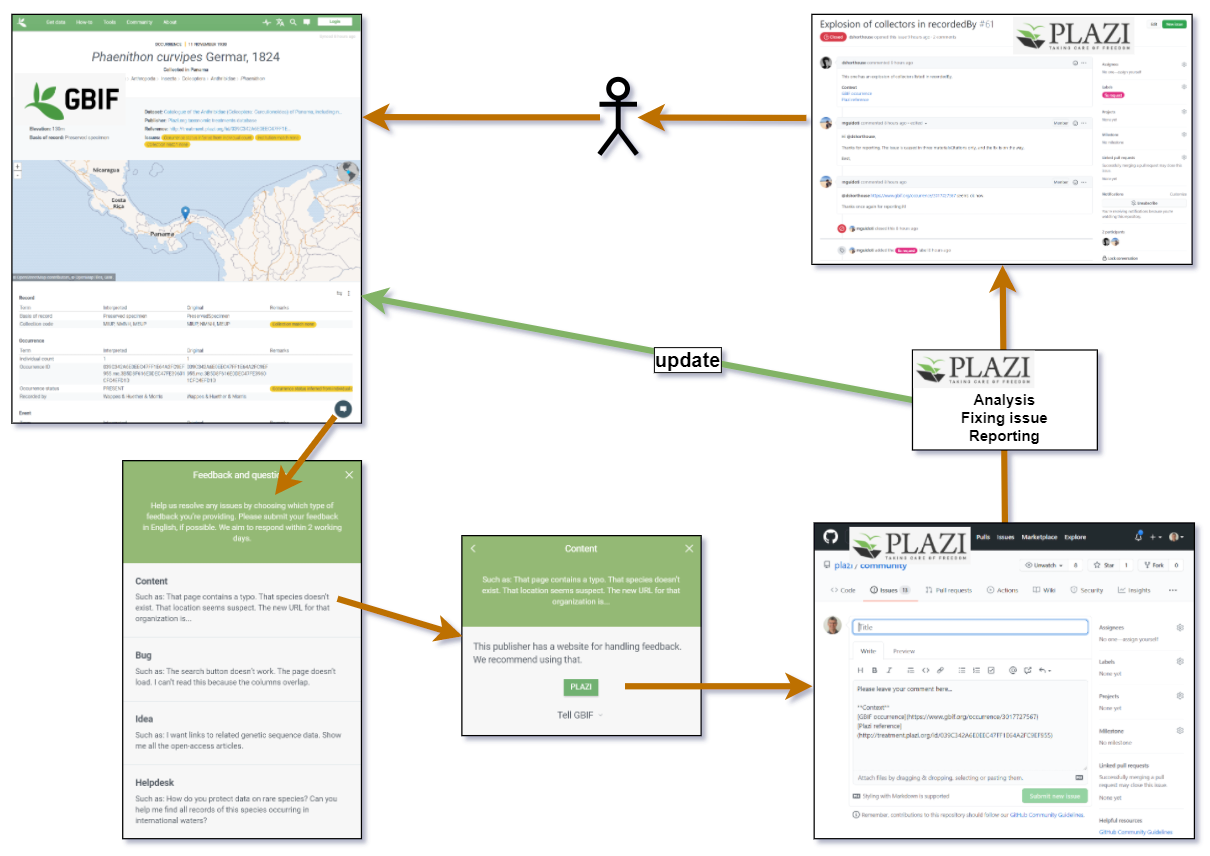
The launch of the new feedback button in GBIF reflects the commitment by GBIF and Plazi to care about the data. It allows a user to send a request for a review of specific data to Plazi. Its care team will respond in a very short time and notify the result, once the issue has been resolved, including the update of the record in GBIF. At the same time, these requests will be collected to analyse the reason for the issues and develop respective measures to mitigate them. With the users input we can improve our production to provide fitter data.
Liberating data is a great challenge from building technologies, operations to funding on the one hand, and on the other hopefully contributes more to a new way scientists publish their research results.
